An NGS instrument (Next-Generation Sequencing instrument) is a laboratory machine used to determine the order of nucleotides in DNA or RNA at extremely high throughput. These systems can sequence millions to billions of DNA fragments simultaneously, allowing researchers to analyze entire genomes, transcriptomes, or microbial communities in a single experiment.
NGS instruments are the core hardware used in genomics laboratories, enabling modern applications such as whole-genome sequencing, cancer genomics, pathogen surveillance, and gene expression analysis.
1. What “Next-Generation Sequencing” Means
DNA is composed of four nucleotide bases:
-
A – Adenine
-
T – Thymine
-
C – Cytosine
-
G – Guanine
Sequencing determines the exact order of these bases along a DNA molecule.
Earlier sequencing technologies (such as Sanger sequencing) could only read one DNA fragment at a time. NGS technologies dramatically improved efficiency by sequencing millions of fragments in parallel, which reduces cost and increases speed.
For perspective:
| Method | Throughput | Cost per genome |
|---|---|---|
| Sanger sequencing | ~1 fragment/run | Very expensive |
| NGS | Millions–billions fragments/run | Much cheaper |
This parallelization is why NGS enabled modern large-scale genomics projects.
2. What an NGS Instrument Actually Does
An NGS instrument performs three main technical functions:
-
Amplify DNA fragments
-
Read nucleotide incorporation events
-
Convert signals into digital sequence data
Inside the machine are several subsystems:
Key Hardware Components
1. Flow Cell
A flow cell is a small glass slide containing microscopic wells or lanes where DNA fragments attach and are sequenced.
Each well holds many identical copies of a DNA fragment.
2. Fluidics System
The instrument pumps reagents across the flow cell:
-
DNA polymerase
-
fluorescent nucleotides
-
buffers
These chemicals enable the sequencing reactions.
3. Imaging System
Most NGS systems detect fluorescent signals emitted when nucleotides are incorporated into DNA strands.
The machine contains:
-
high-resolution cameras
-
lasers
-
optical filters
These capture images of millions of sequencing reactions simultaneously.
4. Computational Hardware
The raw optical signals must be processed into sequence reads. The system therefore includes:
-
onboard processing computers
-
base-calling algorithms
-
data storage and export capability
A single run can produce hundreds of gigabytes of data.
3. The NGS Workflow
Although the instrument performs sequencing itself, NGS experiments involve several steps before and after the machine run.
Step 1: DNA or RNA Extraction
Biological samples are first processed to isolate genetic material from sources such as:
-
blood
-
tissue
-
bacteria
-
environmental samples
-
food products
If RNA is studied, it is converted to complementary DNA (cDNA).
Step 2: Library Preparation
DNA molecules are cut into small fragments (typically 150–500 base pairs).
Special adapter sequences are attached to the ends. These adapters:
-
allow DNA to bind to the flow cell
-
contain barcodes for multiplexing samples
-
provide priming sites for sequencing reactions
The resulting set of fragments is called a sequencing library.
Step 3: Cluster Amplification
Inside the instrument or a separate device, fragments attach to the flow cell and are amplified into clusters.
Each cluster contains thousands of identical DNA molecules derived from a single fragment.
Amplification increases signal strength for detection.
Step 4: Sequencing
The machine then performs sequencing-by-synthesis (in the case of common platforms).
During each cycle:
-
A fluorescently labeled nucleotide is added.
-
DNA polymerase incorporates the correct base.
-
The instrument captures an image to record which nucleotide was added.
-
The fluorescent tag is removed.
-
The next cycle begins.
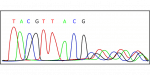
Repeating this cycle produces a sequence read such as:
Step 5: Data Analysis
After sequencing, bioinformatics software processes the reads to:
-
reconstruct genomes
-
identify mutations
-
quantify gene expression
-
classify microorganisms
This step often requires high-performance computing.
4. Major Types of NGS Instruments
Several companies manufacture NGS platforms, each using slightly different chemistry and detection methods.
1. Sequencing-by-Synthesis Platforms
These dominate the market.
Typical characteristics:
-
optical detection
-
fluorescent nucleotides
-
short reads (100–300 bp)
Widely used for:
-
whole genome sequencing
-
RNA sequencing
-
clinical genomics
2. Semiconductor Sequencers
These detect hydrogen ions released during nucleotide incorporation, eliminating the need for fluorescence imaging.
Advantages:
-
faster runs
-
simpler optics
3. Single-Molecule Long-Read Sequencers
Some systems sequence very long DNA fragments (10,000–100,000 bases or more).
Benefits include:
-
better genome assembly
-
detection of structural variants
-
improved analysis of repetitive DNA regions
What Are The Quality metrics Used?
In next-generation sequencing (NGS), quality metrics are critical to assess the reliability, accuracy, and overall success of a sequencing experiment. These metrics help determine whether the data can be confidently used for downstream analyses like variant calling, gene expression profiling, or genome assembly. Here’s a detailed breakdown:
1. Base Quality Metrics
-
Phred Quality Score (Q-score):
Measures the probability that a base is called incorrectly.Q=−10⋅log10(Perror)Q = -10 \cdot \log_{10}(P_{\text{error}})
-
Example: Q30 = 0.1% chance of error (1 in 1000 bases wrong).
-
Higher Q-scores indicate more reliable base calls.
-
-
Mean/Median Base Quality: Average quality across all bases in reads.
-
Per-Base Sequence Quality: Checks if some regions of reads have systematically lower quality.
2. Read-Level Metrics
-
Read Length Distribution: Confirms that reads are of expected length (important for paired-end vs. single-end sequencing).
-
Read Count / Yield: Total number of reads generated.
-
Duplicate Reads: Fraction of reads that are exact duplicates (may indicate PCR bias or low library complexity).
3. Coverage Metrics
-
Depth of Coverage: Number of times a nucleotide is sequenced.
-
Critical for detecting variants accurately.
-
Typical thresholds: 30× for human genome variant calling.
-
-
Uniformity of Coverage: Measures how evenly reads cover the target region or genome.
-
Uneven coverage can bias downstream analyses.
-
-
Breadth of Coverage: Proportion of target bases covered at a minimum depth (e.g., ≥10×).
4. Mapping and Alignment Metrics
-
Alignment Rate: Percentage of reads that successfully map to the reference genome.
-
Uniquely Mapped Reads: Fraction of reads that map to a single location.
-
Mismatch Rate: Average number of mismatches per aligned read.
-
Insert Size Distribution: For paired-end reads, checks expected distance between paired reads.
5. Library/Sequencing Bias Metrics
-
GC Bias: Assesses whether regions with extreme GC content are underrepresented.
-
Adapter Content: Fraction of reads containing adapter sequences that may require trimming.
-
K-mer Analysis: Detects overrepresented sequences or contamination.
6. Error and Contamination Metrics
-
Error Rate: Fraction of bases incorrectly called (estimated from Q-scores or control samples).
-
Contamination Checks: Detect unexpected sequences from other species, viruses, or index hopping in multiplexed runs.
7. Overall Run Quality
-
Q30 Yield: Fraction of bases with Q-score ≥30; a commonly reported metric by Illumina.
-
Cluster Density (Illumina): Optimal density of DNA clusters on the flow cell. Too high → overlapping clusters; too low → low yield.
8. Optional Downstream Metrics
Depending on the experiment, additional quality checks may include:
-
Variant Calling Metrics: Transition/transversion ratio, heterozygosity rate.
-
RNA-seq Metrics: Mapping to exons vs. introns, rRNA contamination.
-
ChIP-seq/ATAC-seq Metrics: Fraction of reads in peaks (FRiP score), signal-to-noise ratio.
Summary
| Metric Type | Key Metrics | Purpose |
|---|---|---|
| Base-level | Phred score, per-base quality | Assess sequencing accuracy |
| Read-level | Read length, duplicates | Library quality, yield |
| Coverage | Depth, breadth, uniformity | Confidence in variant detection |
| Alignment | Mapping rate, mismatch rate | Sequencing and library fidelity |
| Bias | GC bias, adapter content | Detect technical artifacts |
| Error/Contamination | Error rate, cross-species reads | Ensure sample integrity |
| Run-level | Q30, cluster density | Evaluate overall sequencing run |
High-quality NGS data typically shows: high Q30 scores (>80%), uniform coverage, low duplication, low adapter contamination, and high mapping rates (>95%).
5. What NGS Instruments Are Used For
NGS technology has become essential across many scientific and industrial domains.
Medical Genomics
NGS is widely used in healthcare for:
-
cancer mutation profiling
-
rare disease diagnosis
-
prenatal genetic screening
-
pathogen identification
Hospitals and clinical labs run NGS tests to identify genetic variants that influence disease or treatment response.
Pharmaceutical and Biotechnology Research
Drug discovery companies use NGS to:
-
study gene expression
-
identify therapeutic targets
-
analyze CRISPR editing outcomes
-
characterize cell lines
NGS data often informs precision medicine strategies.
Microbiome Research
NGS allows scientists to analyze complex microbial communities.
Examples include:
-
gut microbiome analysis
-
soil microbiology
-
fermentation microbiology
-
environmental monitoring
Instead of culturing microbes, sequencing directly identifies them.
Agriculture and Food Science
NGS helps improve crop and livestock breeding by identifying genes associated with traits such as:
-
drought resistance
-
yield
-
disease resistance
Food companies also use NGS for food safety testing and contamination detection.
Infectious Disease Surveillance
Public health agencies use NGS to track pathogens by sequencing viral or bacterial genomes.
This helps detect:
-
outbreak sources
-
transmission chains
-
emerging variants
6. Why NGS Instruments Are Important
The development of NGS transformed biology for several reasons.
Massive Scale
Millions of DNA fragments can be sequenced simultaneously.
Lower Cost
The cost of sequencing a human genome has dropped from ~$100 million in 2001 to under $1,000 today.
Broad Applications
NGS supports fields including:
-
genomics
-
oncology
-
microbiology
-
evolutionary biology
-
synthetic biology
Data-Driven Biology
Modern life sciences increasingly depend on large genomic datasets, which NGS instruments generate.
7. Typical Users of NGS Instruments
Organizations that operate NGS machines include:
-
genomics research institutes
-
biotechnology companies
-
pharmaceutical companies
-
clinical diagnostic laboratories
-
agricultural genetics labs
-
contract research organizations (CROs)
Because the instruments can cost £100,000–£1 million, many organizations use sequencing service providers instead of owning machines themselves.
An NGS instrument is a high-throughput genomic sequencing machine that reads the nucleotide sequence of DNA or RNA molecules. It works by sequencing millions of DNA fragments in parallel using chemical reactions and optical detection systems. The instrument generates large volumes of genetic data that researchers analyze to study genomes, detect mutations, characterize microbes, and understand biological processes.
NGS technology underpins modern genomics and has become a foundational tool across medicine, biotechnology, agriculture, and environmental science.

Leave a Reply